RAG Any Docs with Fused
TL;DR: With a few Fused UDFs you can turn any public docs site into a fully searchable RAG pipeline — connect data, process it, and expose it as an API.
This example shows how to build a RAG pipeline over Overture Maps documentation using Fused — from ingestion to a queryable API endpoint. The same pattern works for any public docs site.
How it works
The pipeline follows a crawl → embed → search pattern, implemented as four UDFs:
site_to_llmstxt— crawls any docs site via its sitemap and writes a cleanllms.txtto S3docs_ragging— embeds thellms.txtin chunks and stores vectors in a LanceDB table at/mount/lancedb/overture_docs_searching— searches by embedding an incoming question and returning the most relevant chunksai_utils— shared helpers for embedding, LanceDB reads/writes, and agentic tool calling
Step 1: Crawl Any Docs Site into llms.txt
The first challenge is getting clean text out of a docs site. HTML is noisy: navbars, footers, sidebars, and scripts all get in the way. The site_to_llmstxt UDF accepts any docs site URL, crawls every page in parallel via crawl_page.map(), strips boilerplate, and writes a single clean llms.txt to S3.
Show site_to_llmstxt UDF code
@fused.udf(cache_max_age='1d') # Caching for 1d to prevent accidental reruns
def udf(
base_url: str = "https://docs.overturemaps.org/",
site_name: str = "Overture_maps_documentation",
mode: str = "full", # "curated" | "full"
sitemap_path: str = "/sitemap.xml",
max_pages: int = 0, # 0 = no limit, useful for testing
output_dir: str = "s3://fused-asset/demos/llms_txt/",
):
"""Generate an llms.txt from any public docs site with a sitemap."""
import re
import time
import requests
import pandas as pd
from xml.etree import ElementTree
from bs4 import BeautifulSoup
def fetch(url: str, retries: int = 3, timeout: int = 15) -> str | None:
headers = {"User-Agent": "llms-txt-generator/1.0 (sitemap crawler)"}
for attempt in range(retries):
try:
r = requests.get(url, headers=headers, timeout=timeout)
r.raise_for_status()
return r.text
except Exception as e:
if attempt == retries - 1:
print(f" ✗ Failed {url}: {e}")
return None
time.sleep(1.5 ** attempt)
def parse_sitemap(xml_text: str) -> list[str]:
"""Parse a sitemap or sitemap index and return all page URLs."""
try:
root = ElementTree.fromstring(xml_text)
except ElementTree.ParseError:
return []
ns = {"sm": "http://www.sitemaps.org/schemas/sitemap/0.9"}
urls = []
for sitemap_tag in root.findall("sm:sitemap", ns):
loc = sitemap_tag.find("sm:loc", ns)
if loc is not None and loc.text:
sub_xml = fetch(loc.text.strip())
if sub_xml:
urls.extend(parse_sitemap(sub_xml))
for url_tag in root.findall("sm:url", ns):
loc = url_tag.find("sm:loc", ns)
if loc is not None and loc.text:
urls.append(loc.text.strip())
return urls
sitemap_url = base_url.rstrip("/") + sitemap_path
print(f"Fetching sitemap: {sitemap_url}")
sitemap_xml = fetch(sitemap_url)
if not sitemap_xml:
raise RuntimeError(f"Could not fetch sitemap at {sitemap_url}")
all_urls = parse_sitemap(sitemap_xml)
print(f"Found {len(all_urls)} URLs in sitemap")
skip_patterns = re.compile(
r"/(tags?|search|404|sitemap|assets|_|static)/|"
r"\.(xml|json|txt|png|jpg|svg|css|js)$",
re.IGNORECASE,
)
page_urls = [u for u in all_urls if not skip_patterns.search(u)]
page_urls = sorted(set(page_urls))
if max_pages > 0:
page_urls = page_urls[:max_pages]
print(f"Processing {len(page_urls)} doc pages (mode={mode})")
print(f"Crawling {len(page_urls)} pages in parallel...")
results = crawl_page.map(
[{"url": u} for u in page_urls],
engine="local",
max_workers=10,
).df()
pages = results.to_dict("records")
print(f"Successfully extracted {len(pages)} pages")
header = f"# {site_name}\n\n"
if mode == "curated":
lines = [header, "## Pages\n"]
for p in pages:
lines.append(f"- [{p['title']}]({p['url']})")
lines.append(f"\n---\n\nGenerated from {base_url} sitemap. Total pages: {len(pages)}")
output = "\n".join(lines)
else:
sep = "=" * 80
sections = [header, sep + "\n"]
for p in pages:
sections.append(f"## {p['title']}")
sections.append(f"URL: {p['url']}\n")
if p["content"]:
sections.append(p["content"])
sections.append("\n" + sep + "\n")
sections.append(f"\n---\n\nGenerated from {base_url} sitemap. Total pages: {len(pages)}")
output = "\n".join(sections)
if output_dir:
import fsspec
safe_name = re.sub(r"[^\w]+", "_", site_name.lower()).strip("_")
output_path = output_dir.rstrip("/") + f"/{safe_name}/llms.txt"
with fsspec.open(output_path, "w") as f:
f.write(output)
print(f"Written to {output_path}")
return output_path
else:
return output
@fused.udf
def crawl_page(url: str = "https://docs.overturemaps.org/"):
"""Fetch and extract content from a single docs page. Used by udf.map()."""
import re
import time
import requests
from bs4 import BeautifulSoup
headers = {"User-Agent": "llms-txt-generator/1.0 (sitemap crawler)"}
for attempt in range(3):
try:
r = requests.get(url, headers=headers, timeout=15)
r.raise_for_status()
html = r.text
break
except Exception:
if attempt == 2:
return None
time.sleep(1.5 ** attempt)
else:
return None
soup = BeautifulSoup(html, "html.parser")
for tag in soup.select(
"nav, footer, aside, .sidebar, .toc, .table-of-contents, "
".navbar, .pagination, .edit-this-page, script, style, "
"[class*='sidebar'], [class*='navbar'], [class*='footer'], "
"[class*='toc'], [class*='pagination'], [class*='breadcrumb'], "
"[aria-hidden='true']"
):
tag.decompose()
title = ""
h1 = soup.find("h1")
if h1:
title = h1.get_text(strip=True)
if not title:
title_tag = soup.find("title")
if title_tag:
title = re.sub(r"\s*[|–—-].*$", "", title_tag.get_text(strip=True))
content_el = (
soup.find("article")
or soup.find(class_=re.compile(r"md-content|markdown|content|documentation"))
or soup.find("main")
or soup.body
)
if content_el is None:
import pandas as pd
return pd.DataFrame([{"url": url, "title": title, "description": "", "content": ""}])
def extract_content(el) -> str:
from bs4 import NavigableString, Tag
BLOCK_TAGS = {"p", "div", "section", "li", "dt", "dd", "blockquote",
"h1", "h2", "h3", "h4", "h5", "h6", "tr", "thead", "tbody"}
def walk(node) -> str:
if isinstance(node, NavigableString):
return str(node)
if not isinstance(node, Tag):
return ""
tag = node.name.lower() if node.name else ""
if tag == "pre":
inner_code = node.find("code")
lang = ""
if inner_code and inner_code.get("class"):
for cls in inner_code.get("class", []):
m = re.match(r"language-(\w+)", cls)
if m:
lang = m.group(1)
break
code_text = (inner_code or node).get_text()
return f"\n\n```{lang}\n{code_text.strip()}\n```\n\n"
if tag == "code":
text = node.get_text()
if "\n" in text:
return f"\n\n```\n{text.strip()}\n```\n\n"
return f"`{text}`"
if tag in {"h1", "h2", "h3", "h4", "h5", "h6"}:
level = int(tag[1])
return f"\n\n{'#' * level} {node.get_text(strip=True)}\n\n"
parts = [walk(child) for child in node.children]
text = "".join(parts)
if tag in BLOCK_TAGS:
text = text.strip()
return f"\n\n{text}\n\n" if text else ""
return text
raw = walk(el)
return re.sub(r"\n{3,}", "\n\n", raw).strip()
content = extract_content(content_el)
description = ""
for line in content.splitlines():
line = line.strip()
if len(line) >= 40 and not line.startswith("#") and not line.startswith("`"):
description = line[:160] + ("..." if len(line) > 160 else "")
break
import pandas as pd
return pd.DataFrame([{"url": url, "title": title, "description": description, "content": content}])
Step 2: Chunk, Embed, and Store in LanceDB
The docs_ragging UDF fetches the llms.txt from S3, splits it into overlapping chunks, embeds each chunk using the Qwen3-Embedding-8B model via OpenRouter, and writes the result to a LanceDB table on the shared volume at /mount/lancedb/.
Show docs_ragging UDF code
# ===================== CONFIGURATION =====================
LANCEDB_TABLE_NAME = "overture_docs_17_04_2026"
LANCEDB_BASE_PATH = "/mount/lancedb/"
EMBEDDING_MODEL = "qwen/qwen3-embedding-8b"
SOURCE_S3_PATH = "s3://fused-asset/demos/llms_txt/overture_maps_documentation/llms.txt"
CHUNK_SIZE = 1300
OVERLAP_PERCENT = 0.15
OVERLAP_SIZE = int(CHUNK_SIZE * OVERLAP_PERCENT)
EMBEDDING_BATCH_SIZE = 400
# =========================================================
@fused.udf(cache_max_age=0)
def udf(
table_name: str = LANCEDB_TABLE_NAME,
overwrite: bool = True,
):
import pandas as pd
import fsspec
ai = fused.load("ai_utils")
print(f"Fetching docs from: {SOURCE_S3_PATH}")
with fsspec.open(SOURCE_S3_PATH, "r") as f:
content = f.read()
print(f"Fetched {len(content):,} characters")
chunks = chunk_with_overlap(content, CHUNK_SIZE, OVERLAP_SIZE)
print(f"Created {len(chunks)} chunks")
df = pd.DataFrame(
{
"chunk_id": list(range(1, len(chunks) + 1)),
"code": chunks,
"source_url": SOURCE_S3_PATH,
}
)
print("Embedding chunks via ai_utils.embed_df...")
df = ai.embed_df(
df,
text_col="code",
embedding_col="embedding",
provider="qwen",
embedding_model=EMBEDDING_MODEL,
batch_size=EMBEDDING_BATCH_SIZE,
max_workers=16,
)
print(f"Writing {len(df)} rows to LanceDB table: {table_name} at {LANCEDB_BASE_PATH}")
table_path = ai.write_table(
df,
table_name=table_name,
embedding_col="embedding",
base_path=LANCEDB_BASE_PATH,
overwrite=overwrite,
)
visible_tables = ai.list_tables(base_path=LANCEDB_BASE_PATH)
table_exists_now = ai.table_exists(table_name, base_path=LANCEDB_BASE_PATH)
return pd.DataFrame(
{
"status": ["success"],
"table_name": [table_name],
"table_path": [table_path],
"lancedb_base_path": [LANCEDB_BASE_PATH],
"table_exists_after_write": [table_exists_now],
"source_url": [SOURCE_S3_PATH],
"content_chars": [len(content)],
"chunks": [len(chunks)],
"chunk_size": [CHUNK_SIZE],
"overlap_size": [OVERLAP_SIZE],
}
)
def chunk_with_overlap(text, chunk_size, overlap_size):
"""Split text into chunks with fixed overlap."""
chunks = []
start = 0
text_length = len(text)
while start < text_length:
end = start + chunk_size
chunk = text[start:end]
chunks.append(chunk)
start = end - overlap_size
if start + chunk_size >= text_length and start < text_length:
final_chunk = text[start:]
if final_chunk and final_chunk not in chunks and len(final_chunk) > overlap_size:
chunks.append(final_chunk)
break
return chunks
After running, the UDF returns a status table:
status table_name table_path chunks chunk_size overlap_size content_chars
success overture_docs_17_04_2026 /mount/lancedb/overture_docs_17_04_2026 639 1300 195 706121
Embeddings are generated in batches of 400 chunks with up to 16 concurrent workers. Switching to a different embedding model is a one-line change in the EMBEDDING_MODEL constant at the top of the UDF.
Step 3: Semantic Search as a Fused API
The overture_docs_searching UDF accepts a natural-language question, embeds it with the same model, and runs a cosine similarity search against the stored chunks in LanceDB. Because it's a standard Fused UDF, it's immediately available as an HTTP endpoint.
Show overture_docs_searching UDF code
@fused.udf
def udf(
question: str = "Latest release?",
):
"""Searches Overture docs RAG for matching areas in the documentation."""
import pandas as pd
collection_name = "overture_docs_17_04_2026"
limit = 30
base_path = "/mount/lancedb/"
ai = fused.load("ai_utils")
if not ai.table_exists(collection_name, base_path=base_path):
available = ai.list_tables(base_path=base_path)
return pd.DataFrame(
{
"status": ["error"],
"error": [f"LanceDB table '{collection_name}' was not found."],
"base_path": [base_path],
"available_tables": [", ".join(available) if available else "(none)"],
}
)
df = ai.search(question, collection_name, top_k=limit)
if df is None or len(df) == 0:
return pd.DataFrame({"status": ["empty"], "message": ["No vector matches found."]})
cols_to_drop = [c for c in ["_distance", "_relevance_score", "vector", "embedding"] if c in df.columns]
if cols_to_drop:
df = df.drop(columns=cols_to_drop)
return df.head(5)
Asking "Latest release?" returns a ranked table of the most relevant doc chunks:
chunk_id similarity code
215 0.123 ## Overture Maps Engineering Blog ... August 20, 2025 — 2025-08-20.0 release ...
164 0.112 The 2025-08-20.0 release of Overture data and v1.11.0 of the schema are now available ...
214 0.095 April 22 - Exploring our beta release / May 16 - 2024-05-16-beta.0 release notes ...
213 0.092 ### 2026 — January 21 - 2026-01-21 release notes / February 11 - Overture Has ...
165 0.092 The base, buildings, divisions, places, and transportation themes are in GA ...
Each search request re-embeds the question on every call. Wrap the UDF with @fused.cache to cache results for repeated or identical queries and cut latency significantly.
Using This for Any Site
To RAG a different docs site, update these values:
- In
site_to_llmstxt, updatebase_urlon line 2 of the UDF parameters:
base_url: str = "https://docs.myproject.io/", # URL of the docs site to crawl
- In
docs_ragging, update the two constants at the top of the file:
SOURCE_S3_PATH = "s3://your-bucket/myproject/llms.txt" # S3 path where site_to_llmstxt wrote the output
LANCEDB_TABLE_NAME = "myproject_docs_v1" # unique name for this project's vector table
- In
overture_docs_searching, updatecollection_nameto match:
collection_name = "myproject_docs_v1" # must match LANCEDB_TABLE_NAME above
- Large document sets may take time to index — crawling and embedding hundreds of pages runs in parallel but is still I/O and compute-bound
- Re-indexing overwrites the existing LanceDB table and re-embeds all chunks, which incurs additional embedding API costs; only re-index when the source docs change significantly
Connect to Slack
Once the search UDF is live as a Fused API endpoint, you can connect it to Slack so your team can query the docs directly from a channel. The Fused Slack integration lets you wire any UDF endpoint to a Slack bot — no extra infrastructure needed.
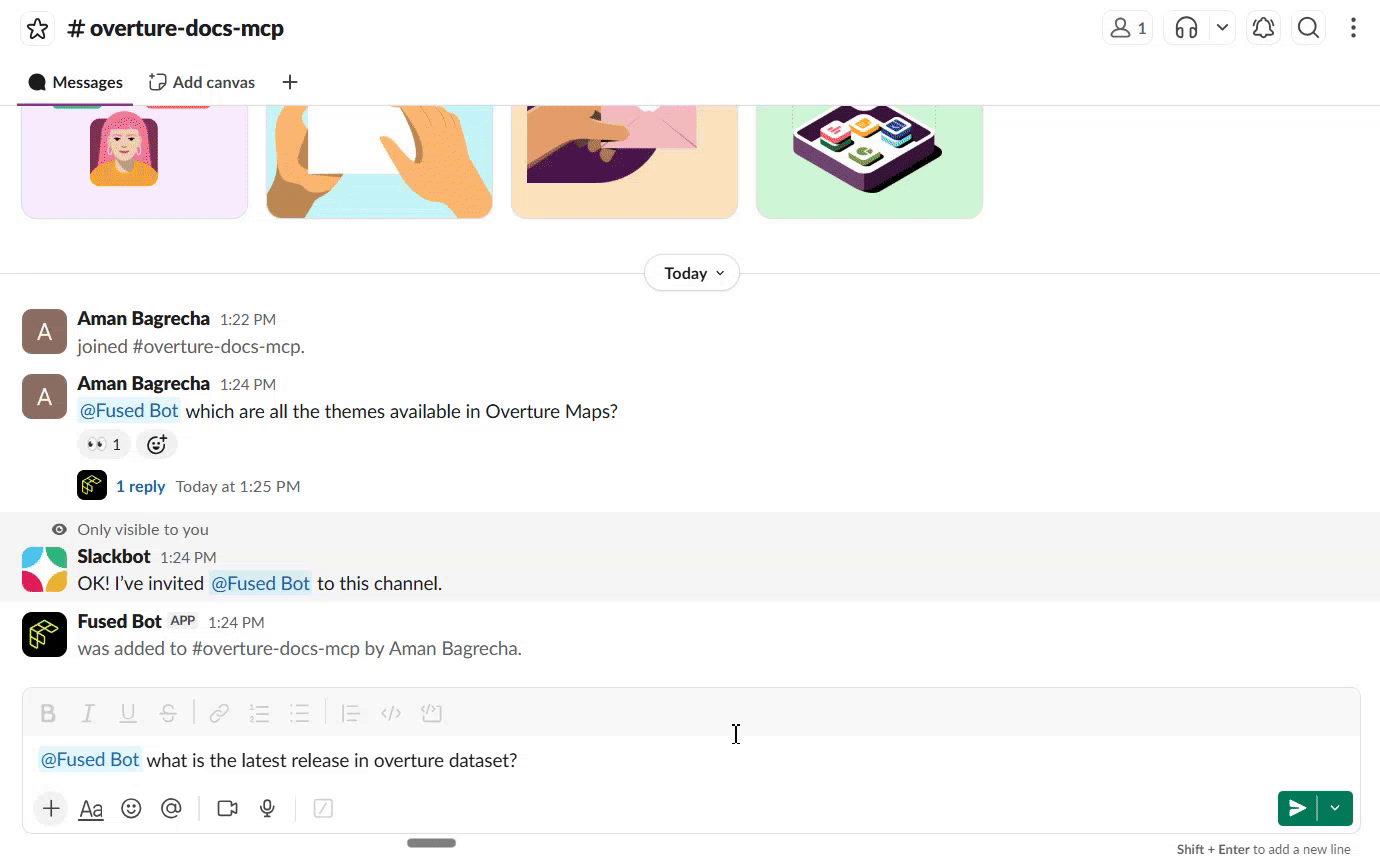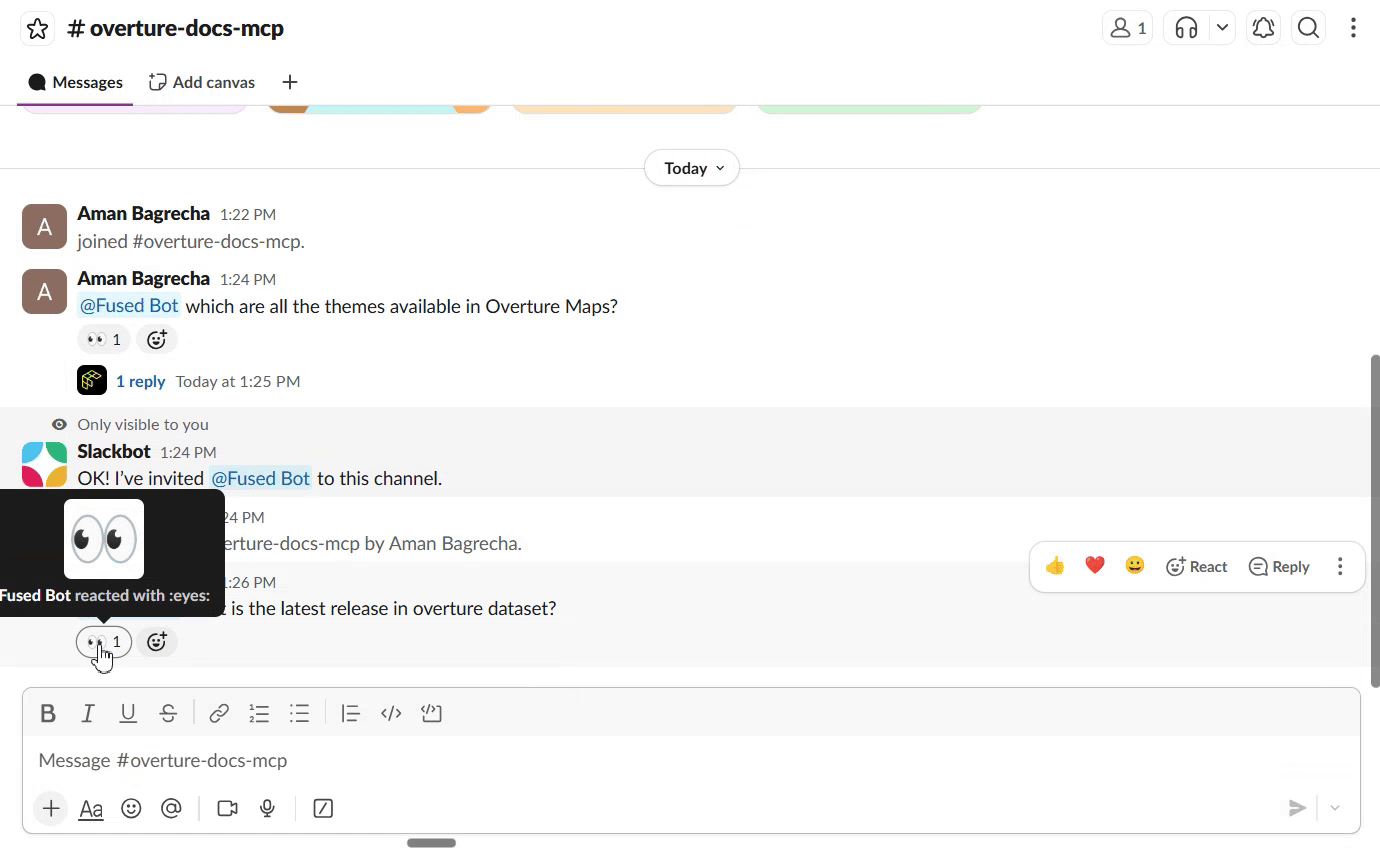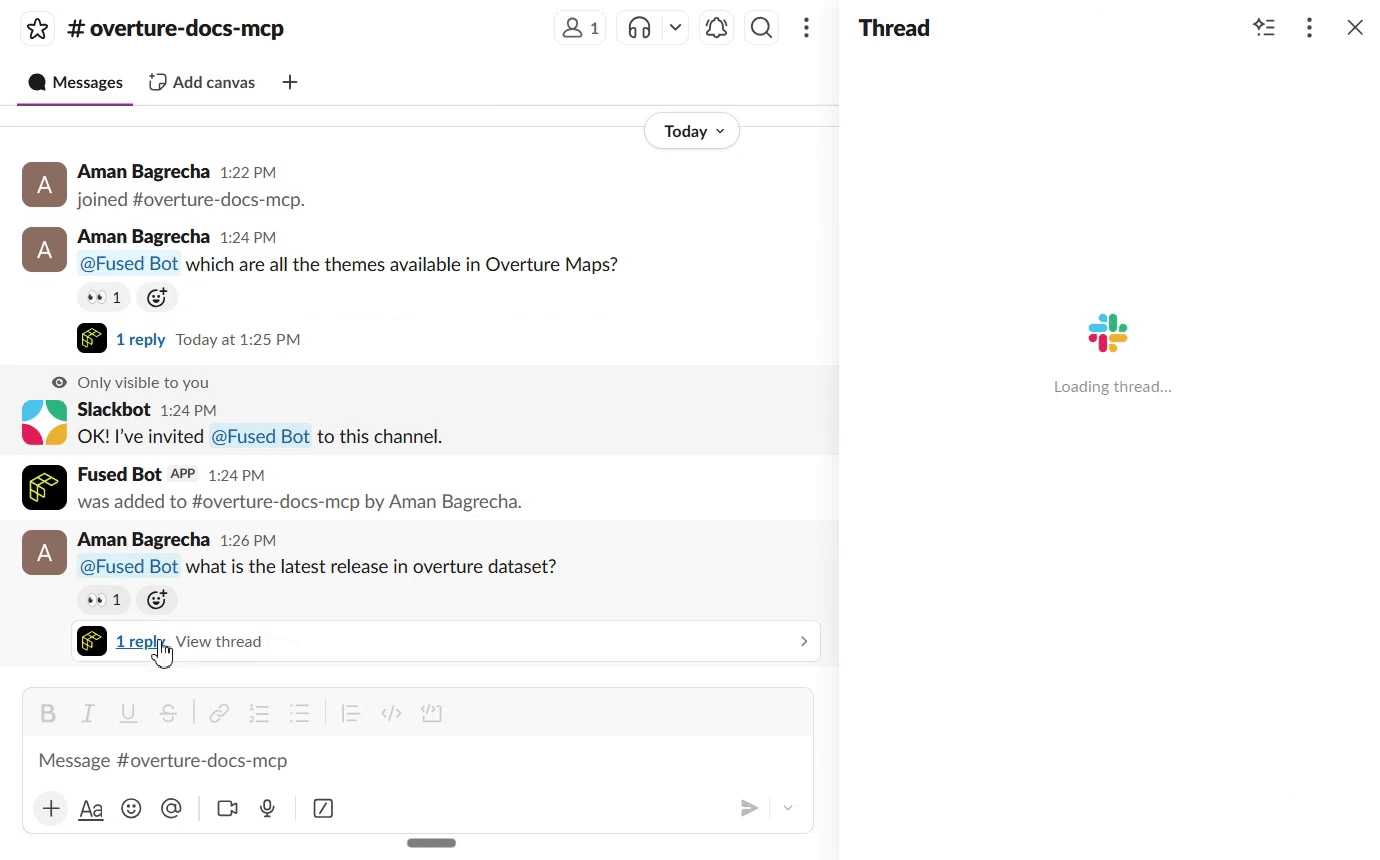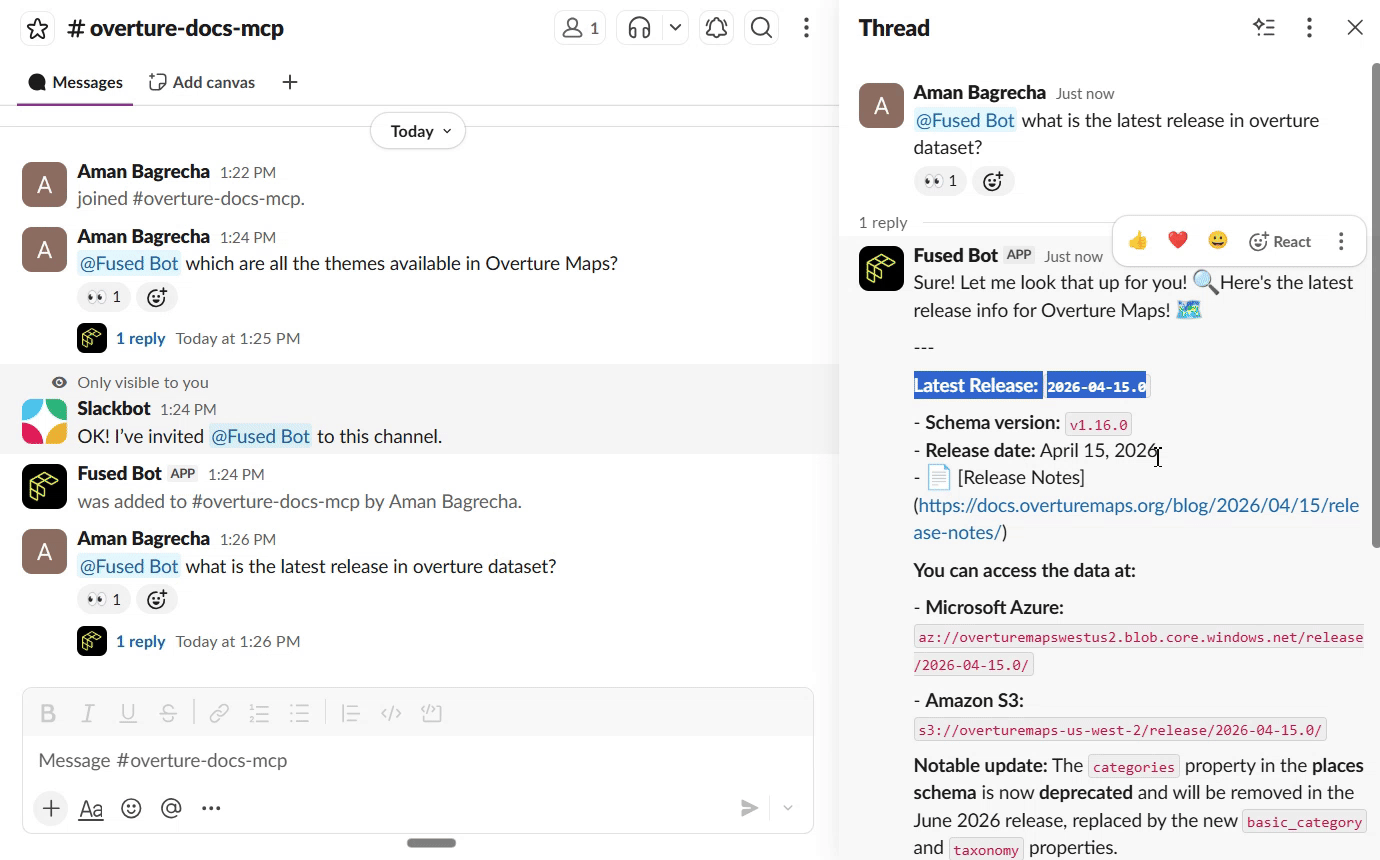
Next Steps
To go further with this pipeline, explore these pages:
- Caching — cache embedding and search results to avoid recomputing on repeated queries
- UDFs as API — expose any UDF as an HTTP endpoint with a shared token
- Slack integration — route search queries through a Slack bot
- Building for Agents — use UDF endpoints as tools in an agentic workflow
Try the full pipeline on Fused Canvas to see all four UDFs wired together.